Can Metagenomic next-generation sequencing give us a headstart against Disease X?
As experts warn of future pandemics, HTX’s Chemical, Biological, Radiological, Nuclear, and Explosives Centre of Expertise is harnessing pathogen-agnostic science to detect unknown and emerging biological threats before they spread widely.
By Cara Yap | Published on 10 March 2026
SCROLL DOWN
Even as we continue to reckon with the human and economic toll of Covid-19, an inescapable question remains: How prepared are we to confront ‘Disease X’? The term, coined by the World Health Organization (WHO), denotes the risk of a future epidemic caused by a pathogen (a disease-causing microbe) not yet known to infect humans.
And it may come sooner than expected, with a growing number of new pathogens – a review published in PubMed Central revealed that virus emergence peaked in the 2000s, comprising 25.5% of new pathogens since the 20th century – and variants caused by factors including human encroachment into wildlife habitats, climate change and globalisation.
And it may come sooner than expected, with a growing number of new pathogens – a review published in PubMed Central revealed that virus emergence peaked in the 2000s, comprising 25.5% of new pathogens since the 20th century – and variants caused by factors including human encroachment into wildlife habitats, climate change and globalisation.
Infographic: HTX/Nicole Lim
But don’t start building your underground bunker just yet. Thanks to modern science, discovering the much dreaded ‘Disease X’ isn’t a shot in the dark.
An advanced technique known as Metagenomic next-generation sequencing (mNGS) allows researchers to scan all the genetic material in a biological sample at once, spotting known pathogens and even completely new microbes. The approach is a form of agnostic testing, which allows scientists to identify unusual or emerging biological threats without prior knowledge of them. Anna Lim, Deputy Director at HTX’s Chemical, Biological, Radiological, Nuclear, and Explosives (CBRNE) Centre of Expertise (CoE), explained that mNGS overcomes certain limitations of traditional biological threat detection methods, which are designed to look for specific, pre-identified pathogens and genetic signatures, and may thus miss highly mutated or engineered variants.
Anna Lim, Deputy Director at HTX’s Chemical, Biological, Radiological, Nuclear, and Explosives (CBRNE) Centre of Expertise (CoE), explained that mNGS overcomes certain limitations of traditional biological threat detection methods, which are designed to look for specific, pre-identified pathogens and genetic signatures, and may thus miss highly mutated or engineered variants.
Unlike conventional methods, mNGS reads millions to billions of DNA or RNA fragments simultaneously – a method known as high-throughput sequencing – producing a comprehensive snapshot of the sample’s biological profile in a single run.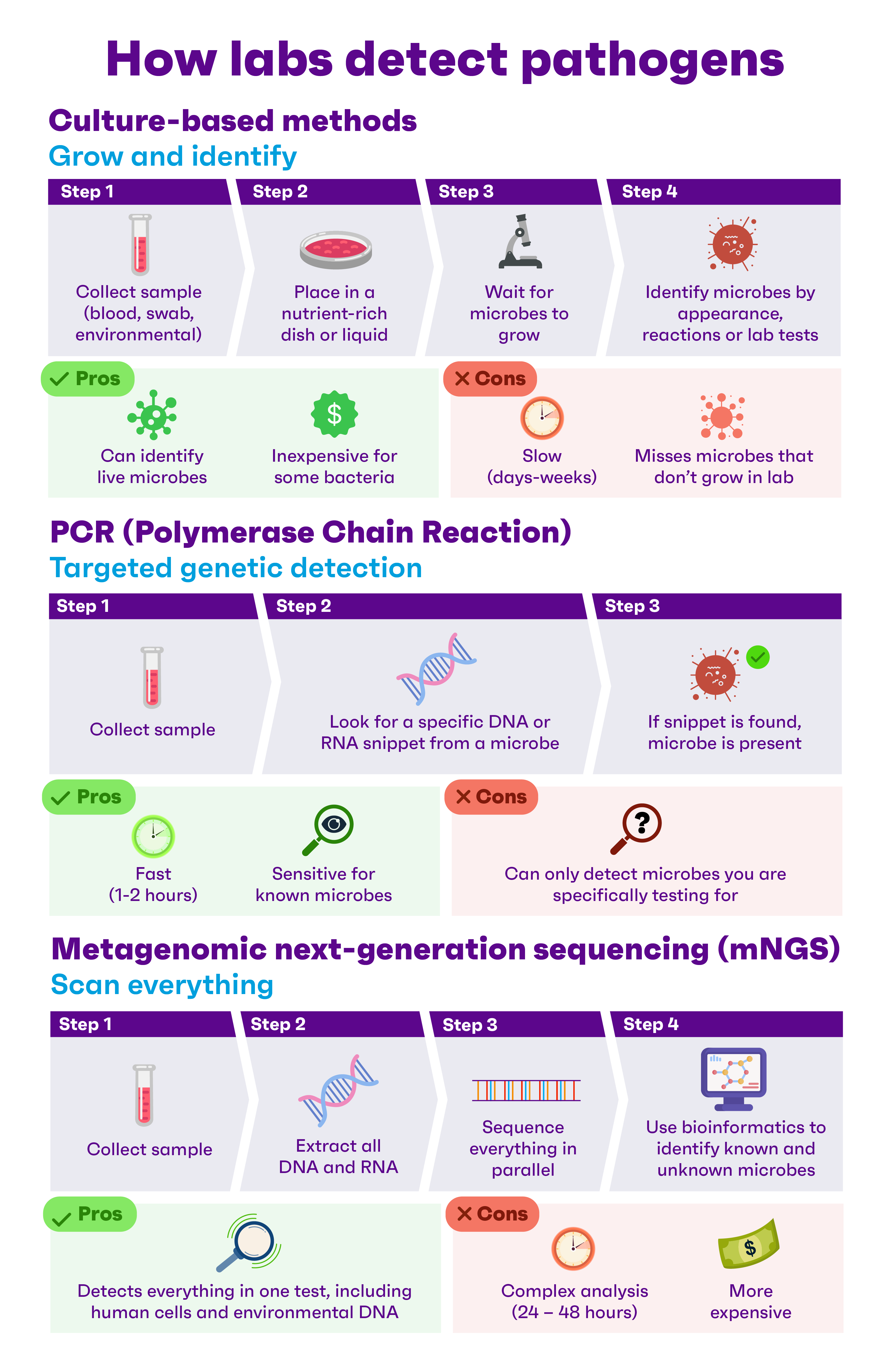
“mNGS shifts detection from confirmation of known biothreats to discovery and early warning,” said Anna. “These include emerging zoonotic pathogens that are poorly genetically characterised, pathogens originating from animal reservoirs, novel and highly mutated species or unusual microbial patterns in the environment.”
Early detection gives researchers and public health authorities a headstart by enabling them to study infectivity (how easily a pathogen spreads from one host to another), virulence (how severe or damaging the infection is) and potential treatments before the biothreat becomes a widespread outbreak.
An advanced technique known as Metagenomic next-generation sequencing (mNGS) allows researchers to scan all the genetic material in a biological sample at once, spotting known pathogens and even completely new microbes. The approach is a form of agnostic testing, which allows scientists to identify unusual or emerging biological threats without prior knowledge of them.
 Anna Lim, Deputy Director at HTX’s Chemical, Biological, Radiological, Nuclear, and Explosives (CBRNE) Centre of Expertise (CoE), explained that mNGS overcomes certain limitations of traditional biological threat detection methods, which are designed to look for specific, pre-identified pathogens and genetic signatures, and may thus miss highly mutated or engineered variants.
Anna Lim, Deputy Director at HTX’s Chemical, Biological, Radiological, Nuclear, and Explosives (CBRNE) Centre of Expertise (CoE), explained that mNGS overcomes certain limitations of traditional biological threat detection methods, which are designed to look for specific, pre-identified pathogens and genetic signatures, and may thus miss highly mutated or engineered variants. Unlike conventional methods, mNGS reads millions to billions of DNA or RNA fragments simultaneously – a method known as high-throughput sequencing – producing a comprehensive snapshot of the sample’s biological profile in a single run.

Infographic: HTX/Nicole Lim
“mNGS shifts detection from confirmation of known biothreats to discovery and early warning,” said Anna. “These include emerging zoonotic pathogens that are poorly genetically characterised, pathogens originating from animal reservoirs, novel and highly mutated species or unusual microbial patterns in the environment.”
Early detection gives researchers and public health authorities a headstart by enabling them to study infectivity (how easily a pathogen spreads from one host to another), virulence (how severe or damaging the infection is) and potential treatments before the biothreat becomes a widespread outbreak.
Why is mNGS more important than ever?
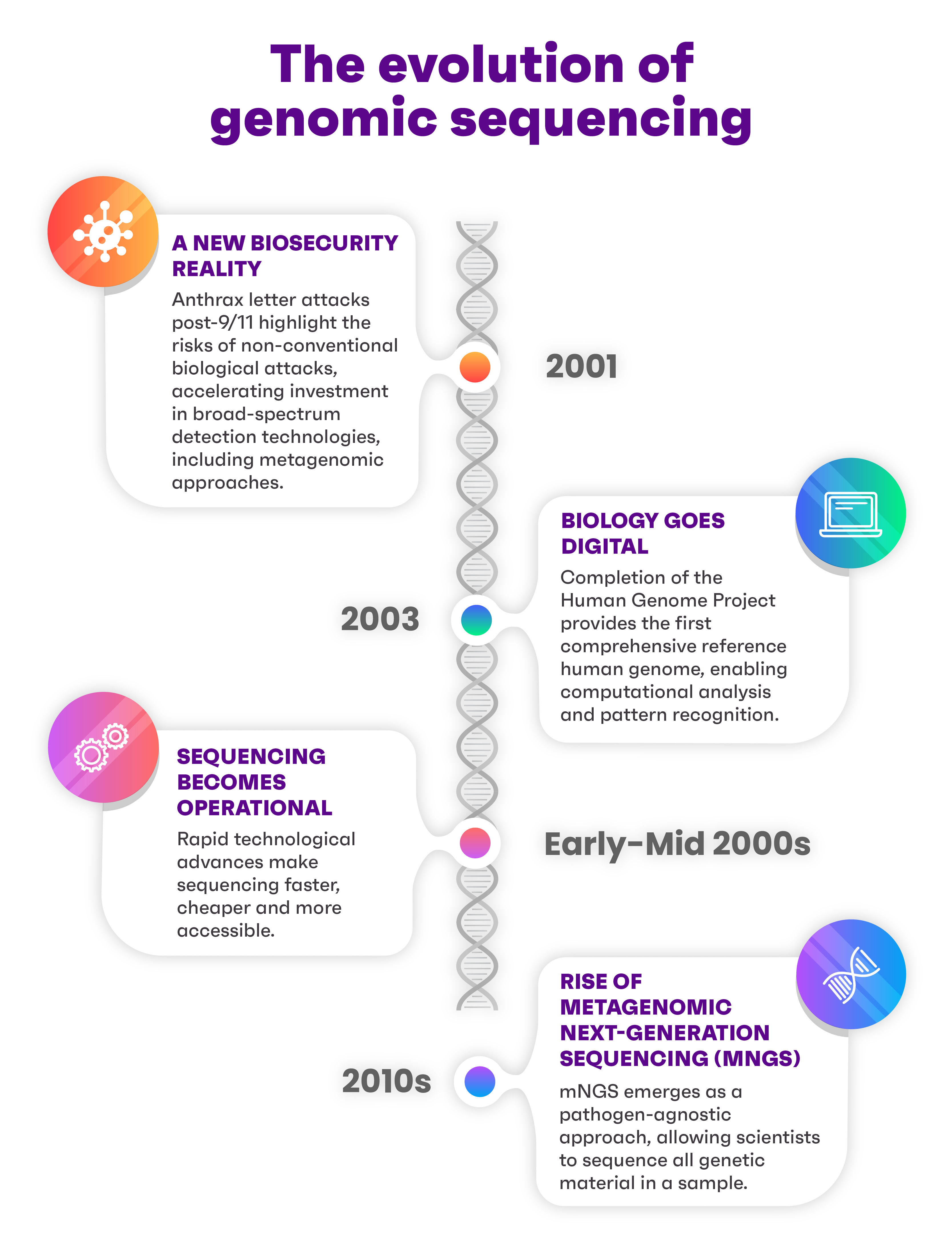
Infographic: HTX/Nicole Lim
mNGS doesn’t just spot pathogens – it can also detect manmade toxins and unusual biological patterns in the environment that may signal a threat, such as a sudden spike in antibiotic-resistance genes.
According to Prof Niranjan Nagarajan, Associate Professor in the School of Medicine and Department of Computer Science at the National University of Singapore and Associate Director and Senior Group Leader at A*STAR Genome Institute of Singapore (A*STAR GIS), Singapore is rapidly advancing in the field of mNGS, spurred by advances in high-throughput sequencing. “As sequencing and analysis platforms mature, the next major milestone would be the widespread development of fully automated systems to execute these workflows safely, routinely and robustly with diverse sample types.”
HTX’s role in tackling biothreats
Bolstering Singapore’s pandemic preparedness and response to biothreat capabilities is CBRNE@Changi, HTX’s latest R&D facility near Changi Airport.
“The facility tackles unknown biothreats by combining pathogen agnostic science with strategic location and operational integration to close the gap between detection and response,” said Anna.
Among the initiatives launched by CBRNE is Project MetaBorders, a collaboration between HTX and A*STAR GIS that aims to establish a baseline profile of the microbiome (the community of microbes that naturally live in an environment) at Singapore’s borders through environmental sampling. The goal is to identify unusual or emerging biological signals early.
“We routinely swab high-touch surfaces across land, sea and air checkpoints – everything from fingerprint scanners to toilets – and also conduct air and subsurface sampling,” said Anna.

“The facility tackles unknown biothreats by combining pathogen agnostic science with strategic location and operational integration to close the gap between detection and response,” said Anna.
Among the initiatives launched by CBRNE is Project MetaBorders, a collaboration between HTX and A*STAR GIS that aims to establish a baseline profile of the microbiome (the community of microbes that naturally live in an environment) at Singapore’s borders through environmental sampling. The goal is to identify unusual or emerging biological signals early.
“We routinely swab high-touch surfaces across land, sea and air checkpoints – everything from fingerprint scanners to toilets – and also conduct air and subsurface sampling,” said Anna.

Infographic: HTX/Nicole Lim
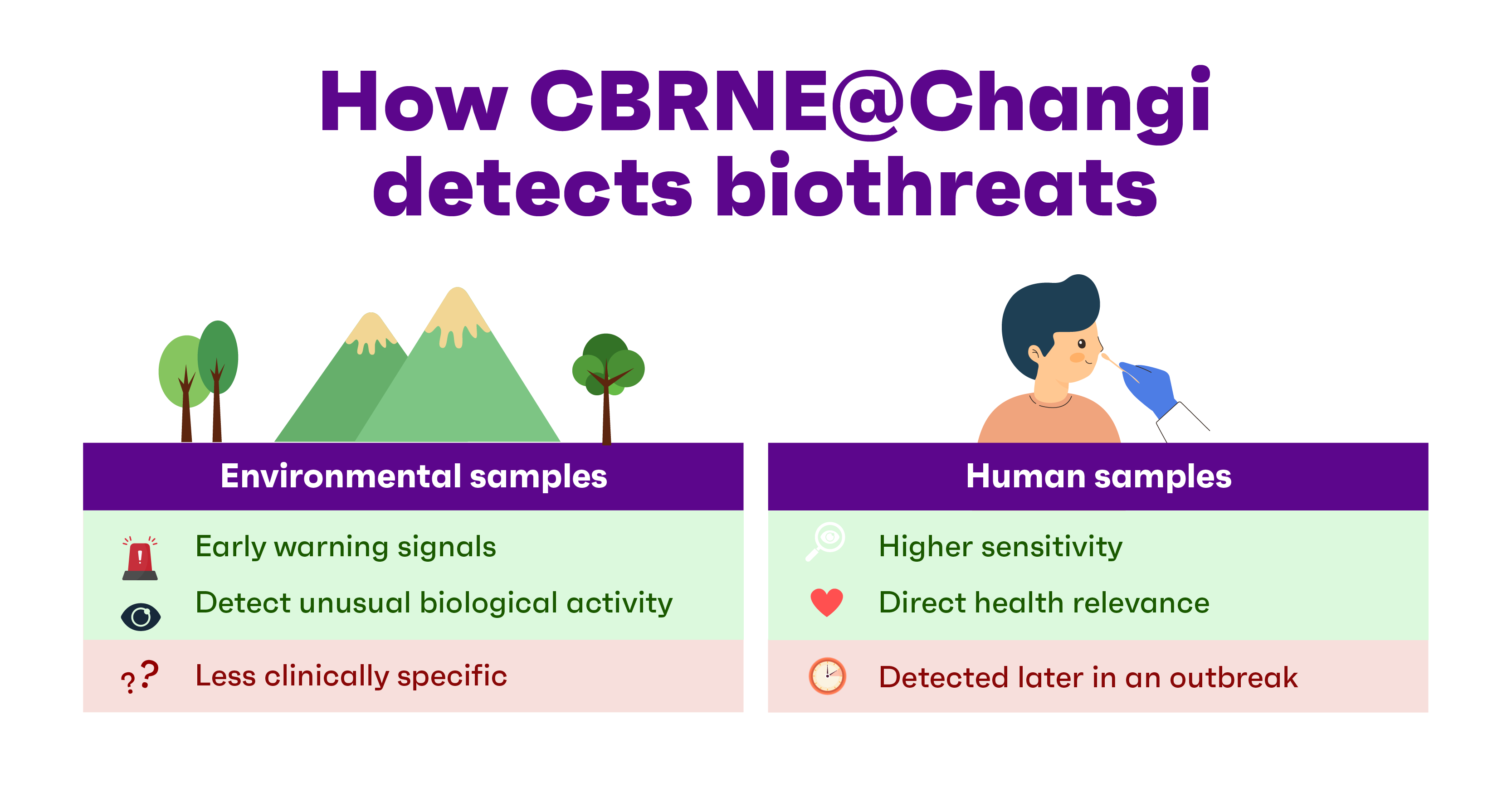
CBRNE@Changi is equipped with advanced mNGS platforms that analyse both environmental and human samples. Here, fully automated DNA/RNA extraction is available to ensure greater consistency, efficiency and traceability, alongside reduced cross-contamination risk.
The shift towards agnostic testing became more pronounced after Covid-19, which underscored the importance of environmental sampling at borders to aid early detection.

Meanwhile, mNGS is becoming a more powerful tool for managing the next pandemic, with AI-assisted and automated interpretation of sequencing results speeding up detection, alongside a more diverse, broader reference database.
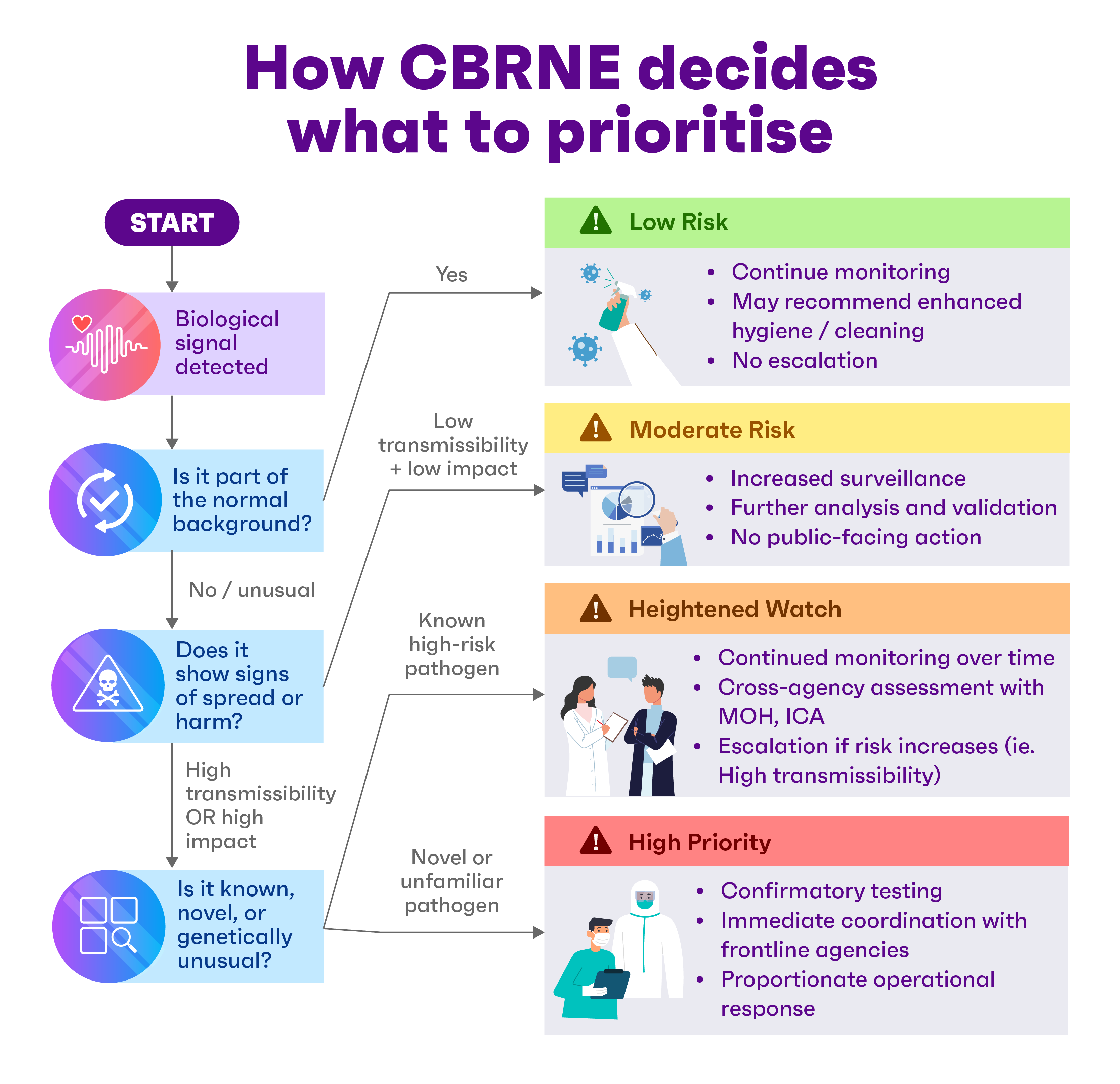
Anna anticipates a future whereby such advanced testing and analysis aren’t confined to central laboratories. Instead, more of it can happen closer to sample collection points – such as at airports, ports or field sites – to enable swifter decisions.


Infographic: HTX/Nicole Lim
Meanwhile, mNGS is becoming a more powerful tool for managing the next pandemic, with AI-assisted and automated interpretation of sequencing results speeding up detection, alongside a more diverse, broader reference database.

Infographic: HTX/Nicole Lim
Anna anticipates a future whereby such advanced testing and analysis aren’t confined to central laboratories. Instead, more of it can happen closer to sample collection points – such as at airports, ports or field sites – to enable swifter decisions.

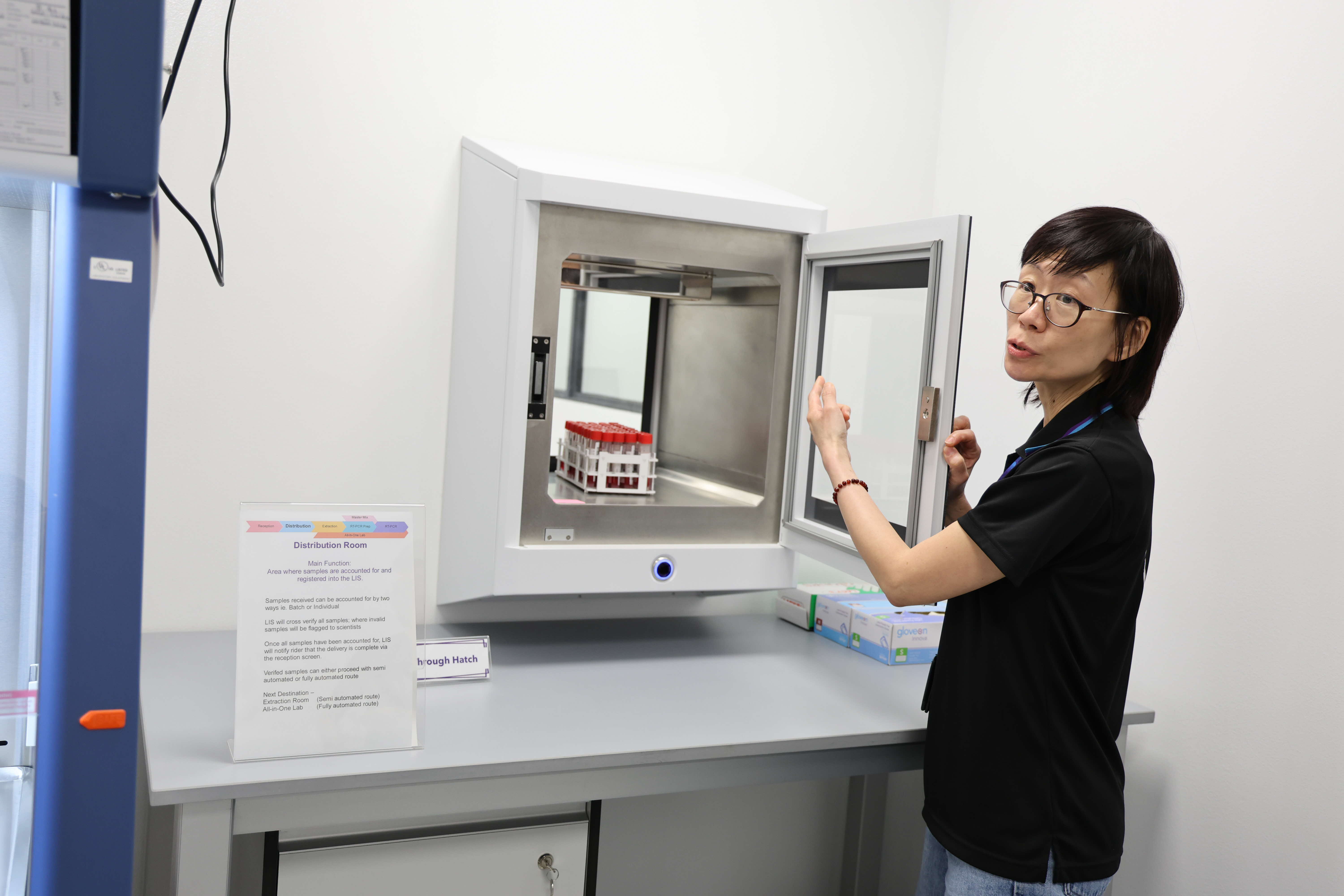
Anna Lim, Deputy Director of HTX’s CBRNE CoE, demonstrates workflow processes at CBRNE@Changi. (Photo: HTX/Dorcas Yang)
And HTX will be right there ahead of the curve. “Success means that metagenomic monitoring at Singapore’s borders is a well-established and routine capability that detects biological change early, reduces uncertainty and enables risk-tiered response without disrupting border operations,” concluded Anna.
